Introduction
Version 2.0 of the synthdid package introduces a modern
formula-based interface similar to lm(),
plm(), and glm(). This vignette demonstrates
how to use the new interface while maintaining full backward
compatibility with existing code.
Why a New Interface?
The new formula interface offers several advantages:
- More intuitive: Follows standard R modeling conventions
- Less code: Single function call instead of multiple steps
-
Better integration: Works with standard R tools
(
summary(),coef(),confint()) -
Easier comparison: Switch between estimators with
update() - Self-documenting: Formula syntax is clearer than matrix indexing
Backward Compatibility
All existing code continues to work! The original matrix-based interface is fully supported:
# Old interface (still works)
data("california_prop99")
setup <- panel.matrices(california_prop99)
old_result <- synthdid_estimate(setup$Y, setup$N0, setup$T0)
old_result
#> synthdid: -15.604 +- NA. Effective N0/N0 = 16.4/38~0.4. Effective T0/T0 = 2.8/19~0.1. N1,T1 = 1,12. [NOT CONVERGED]Basic Usage
Simple Estimation
The new interface uses a formula to specify the outcome and treatment variables:
# New formula interface
result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"))
result
#> Synthetic Difference-in-Differences Estimate
#>
#> Call:
#> synthdid(formula = PacksPerCapita ~ treated, data = california_prop99,
#> index = c("State", "Year"))
#>
#> Treatment Effect: -15.6
#>
#> Units: 38 control, 1 treated
#> Time Periods: 19 pre-treatment, 12 post-treatment
#>
#> Convergence: NOT CONVERGED - lambda (10000/10000 iters), omega (10000/10000 iters)
#> Consider increasing max.iter. Use summary() for details.The formula PacksPerCapita ~ treated specifies: -
Left side: Outcome variable
(PacksPerCapita) - Right side: Treatment
indicator (treated)
The index argument specifies the panel structure: -
First element: Unit identifier (State) - Second element:
Time identifier (Year)
Standard R Methods
The new interface returns objects that work with standard R functions:
# Extract treatment effect
coef(result)
#> treated
#> -15.60379
# Get confidence intervals (requires SE computation)
# confint(result) # Would compute SE automatically
# Summary statistics
summary(result, fast = TRUE) # Use fast=TRUE for jackknife SE
#> Call:
#> synthdid(formula = PacksPerCapita ~ treated, data = california_prop99,
#> index = c("State", "Year"))
#>
#> Treatment Effect Estimate:
#> Estimate
#> treated -15.6 NA NA NA
#>
#> Dimensions:
#> Value
#> Treated units: 1.000
#> Control units: 38.000
#> Effective controls: 16.388
#> Post-treatment periods: 12.000
#> Pre-treatment periods: 19.000
#> Effective periods: 2.784
#>
#> Top Control Units (omega weights):
#> Weight
#> Nevada 0.124
#> New Hampshire 0.105
#> Connecticut 0.078
#> Delaware 0.070
#> Colorado 0.058
#>
#> Top Time Periods (lambda weights):
#> Weight
#> 1988 0.427
#> 1986 0.366
#> 1987 0.206
#>
#> Convergence Status:
#> Overall: NOT CONVERGED
#> Lambda: ✗ (10000/10000 iterations, 100.0% utilization)
#> Omega: ✗ (10000/10000 iterations, 100.0% utilization)
#>
#> Recommendation: Consider increasing max.iter or relaxing min.decrease.
#> Use synthdid_convergence_info() for detailed diagnostics.Comparing Estimators
One of the most powerful features is the ability to easily compare different panel data estimators.
Method Selection
You can choose the estimator at estimation time:
# Synthetic Difference-in-Differences (default)
synthdid_est <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
method = "synthdid")
# Pure Difference-in-Differences
did_est <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
method = "did")
# Synthetic Control
sc_est <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
method = "sc")Updating Models
The update() function allows you to modify an existing
model:
Side-by-Side Comparison
# Compare all three estimators
comparison <- data.frame(
Method = c("SynthDID", "DID", "SC"),
Estimate = c(coef(synthdid_est), coef(did_est), coef(sc_est))
)
print(comparison)
#> Method Estimate
#> 1 SynthDID -15.60379
#> 2 DID -27.34911
#> 3 SC -19.61966The estimates differ because each method makes different assumptions: - SynthDID: Weights both units and time periods - DID: Equal weights on all control units and time periods - SC: Weights only control units (equal time weights)
Standard Errors
Computing Standard Errors
You can request standard errors at estimation time:
# Bootstrap SE (most reliable, but slow)
result_boot <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
se = TRUE,
se_method = "bootstrap",
se_replications = 200)
# Jackknife SE (faster)
result_jack <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
se = TRUE,
se_method = "jackknife")
# Placebo SE (for single treated unit)
result_placebo <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
se = TRUE,
se_method = "placebo",
se_replications = 100)Parallel Processing
For faster SE computation with bootstrap or placebo methods, use parallel processing:
library(future)
# Set up parallel processing
plan(multisession, workers = 4)
# Estimate with parallel SE computation
result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
se = TRUE,
se_method = "bootstrap",
se_replications = 200)
# Reset to sequential
plan(sequential)For a comprehensive guide on parallel processing,
including performance comparisons, best practices, and when to use
sequential vs parallel, see
vignette("parallel-processing").
Thread Management
The package automatically manages BLAS threads to prevent thread oversubscription when using parallel workers. Without proper management:
- Bad scenario: 4 parallel workers × 8 BLAS threads = 32 threads on a 4-core machine
- Performance: Severe degradation due to context switching
The package automatically: 1. Detects when parallel processing is active 2. Sets BLAS to single-threaded mode for each worker 3. Restores BLAS threads after completion
Result: 4 parallel workers × 1 BLAS thread = 4 threads (optimal for 4 cores)
For best thread control, install the optional
RhpcBLASctl package:
install.packages("RhpcBLASctl")This provides reliable control over OpenBLAS and MKL thread counts. Without it, the package falls back to environment variables which may require an R restart to take effect.
Predictions and Diagnostics
Treatment Effect by Period
The predict() function extracts the treatment effect for
each post-treatment period:
# Effect curve
effect_curve <- predict(result, type = "effect")
print(effect_curve)
#> [1] -4.844930 -4.325765 -8.653504 -8.419076 -12.545421 -16.106184
#> [7] -18.905726 -19.350095 -20.883473 -22.781531 -25.944883 -24.484840
# Plot effect curve
plot(effect_curve, type = "b",
xlab = "Post-Treatment Period",
ylab = "Treatment Effect",
main = "Treatment Effect Over Time")
abline(h = 0, lty = 2, col = "gray")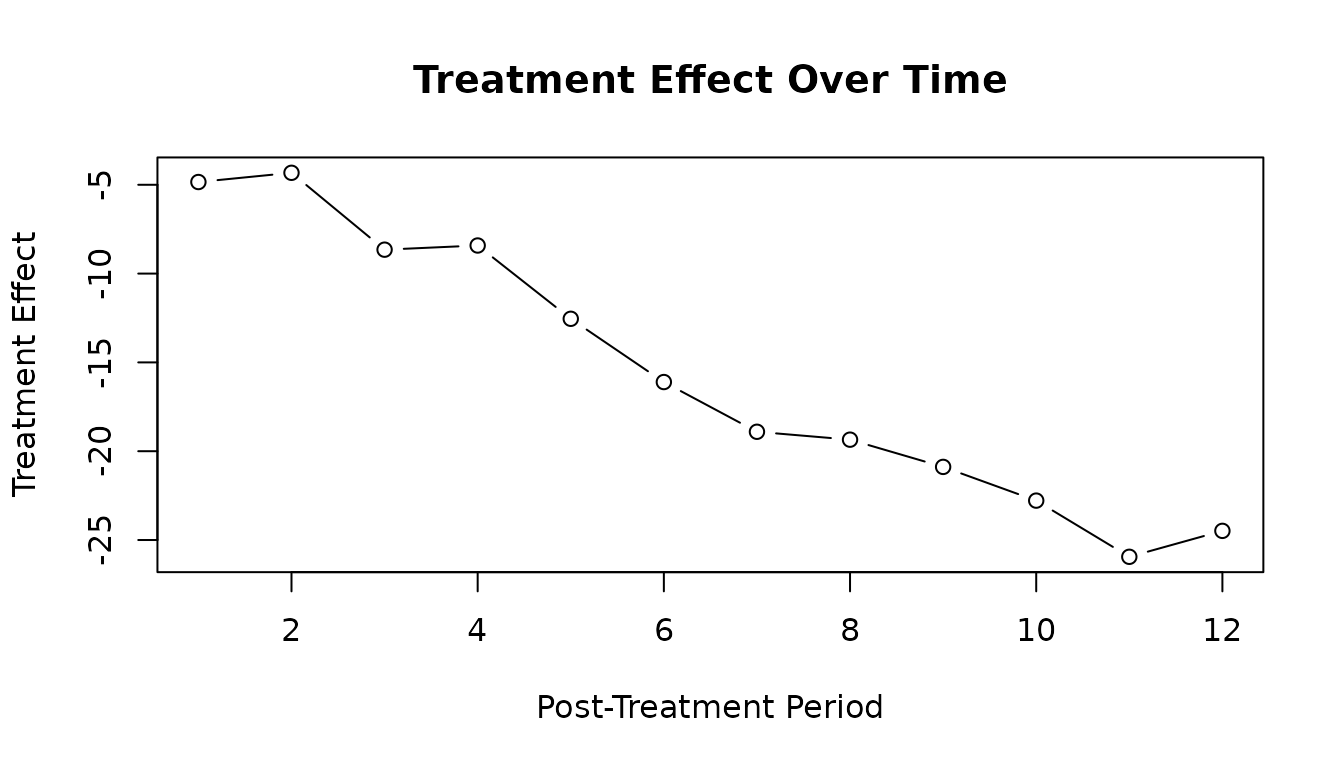
Counterfactual Predictions
What would have happened to California without Prop 99?
# Counterfactual outcomes for treated units
counterfactual <- predict(result, type = "counterfactual")
# Show first 5 periods
counterfactual[, 1:5]
#> 1970 1971 1972 1973 1974
#> 141.8860 145.2028 149.8275 149.1232 150.1937Actual Treated Outcomes
# Actual observed outcomes for treated units
treated_outcomes <- predict(result, type = "treated")
# Show first 5 periods
treated_outcomes[, 1:5]
#> 1970 1971 1972 1973 1974
#> 123.0 121.0 123.5 124.4 126.7Residuals
Extract residuals from different parts of the model:
# Control unit residuals
resid_control <- residuals(result, type = "control")
cat("Control residuals dimension:", dim(resid_control), "\n")
#> Control residuals dimension: 38 31
cat("Mean absolute residual:", mean(abs(resid_control)), "\n")
#> Mean absolute residual: 22.72082
# Pre-treatment period residuals
resid_pretreat <- residuals(result, type = "pretreatment")
cat("Pre-treatment residuals dimension:", dim(resid_pretreat), "\n")
#> Pre-treatment residuals dimension: 39 19
# All residuals
resid_all <- residuals(result, type = "all")
cat("All residuals dimension:", dim(resid_all), "\n")
#> All residuals dimension: 39 31Covariate Adjustment
The formula interface supports time-varying covariates using the
| operator:
# With covariates (if available in your data)
result_cov <- synthdid(PacksPerCapita ~ treated | log_income + unemployment,
data = your_data,
index = c("State", "Year"))The formula
outcome ~ treatment | covariate1 + covariate2 specifies: -
Left of ~: Outcome variable - Right of ~, left
of |: Treatment indicator - Right of |:
Covariates to adjust for
Plotting
All existing plotting functions work with the new interface:
# Standard synthdid plot
plot(result)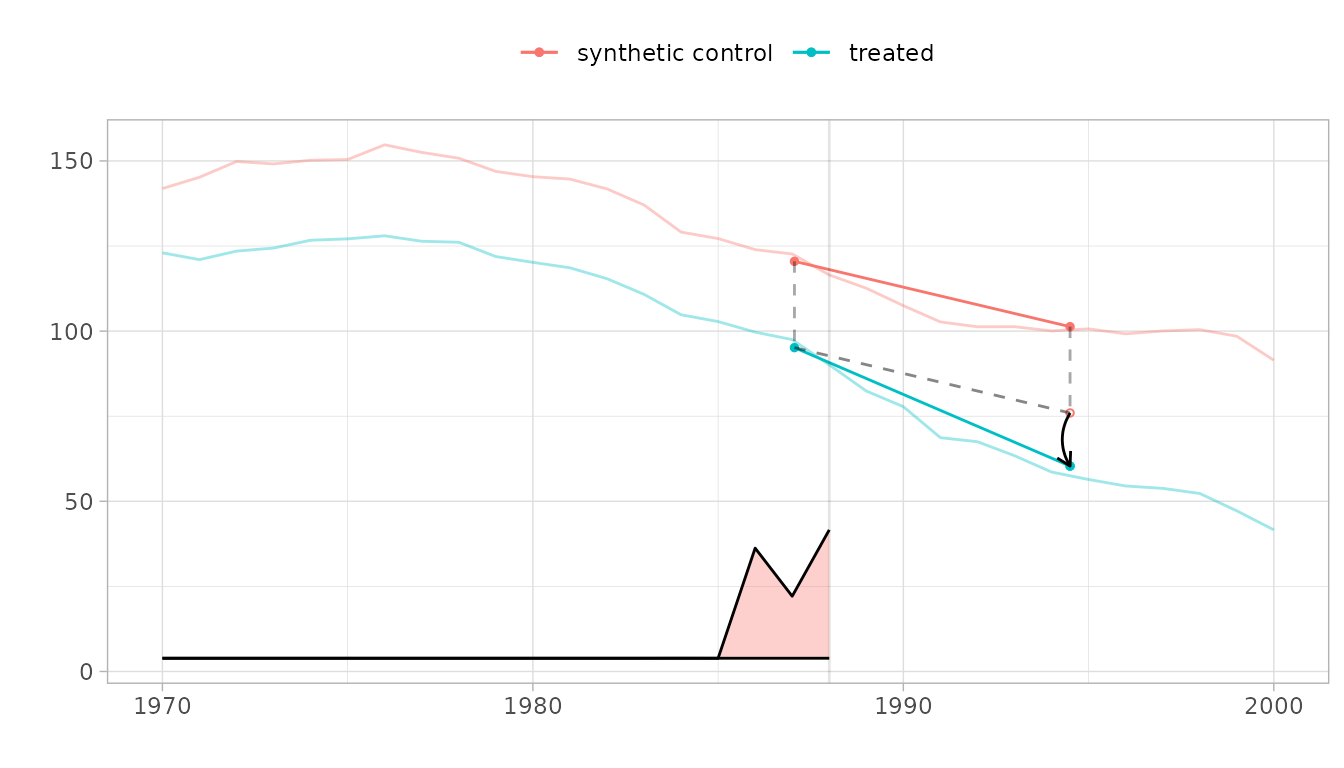
# Control unit contribution plot
synthdid_units_plot(result)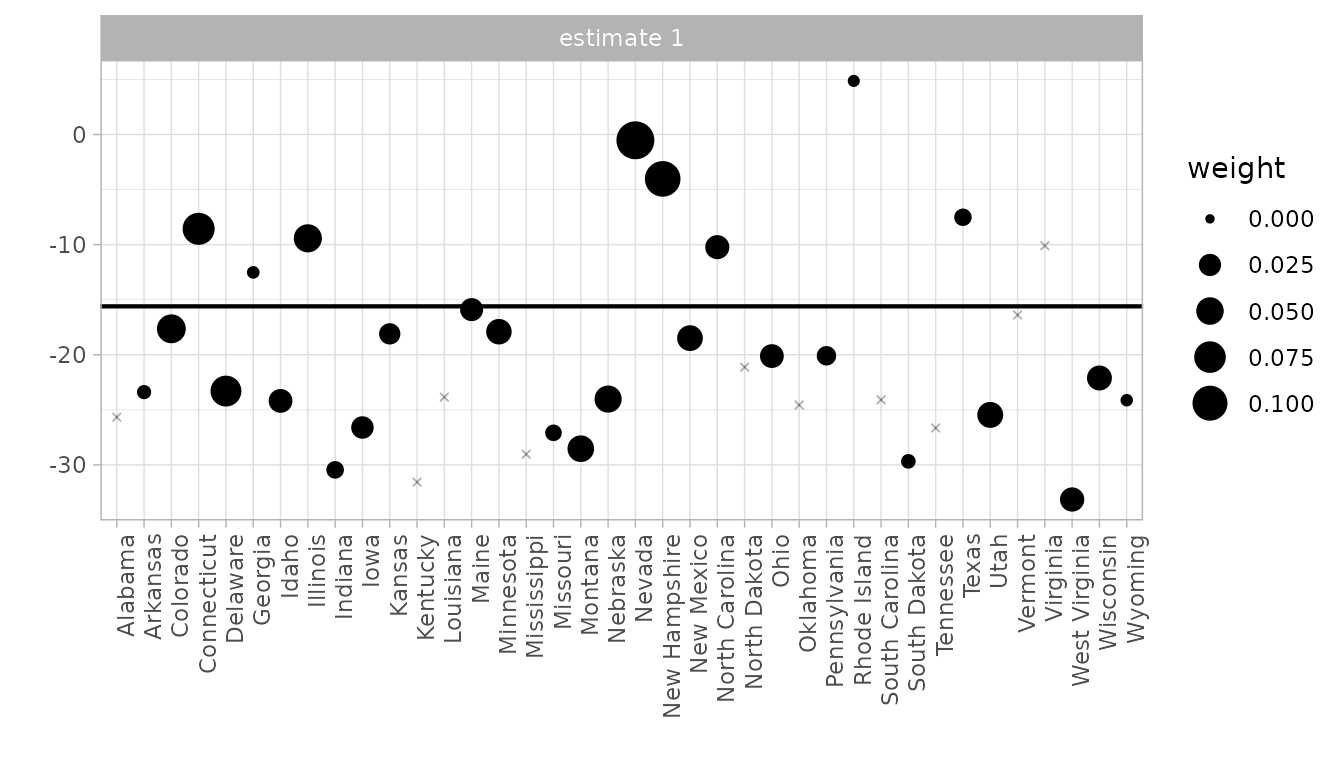
# Check pre-treatment parallel trends
plot(result, overlay = 1)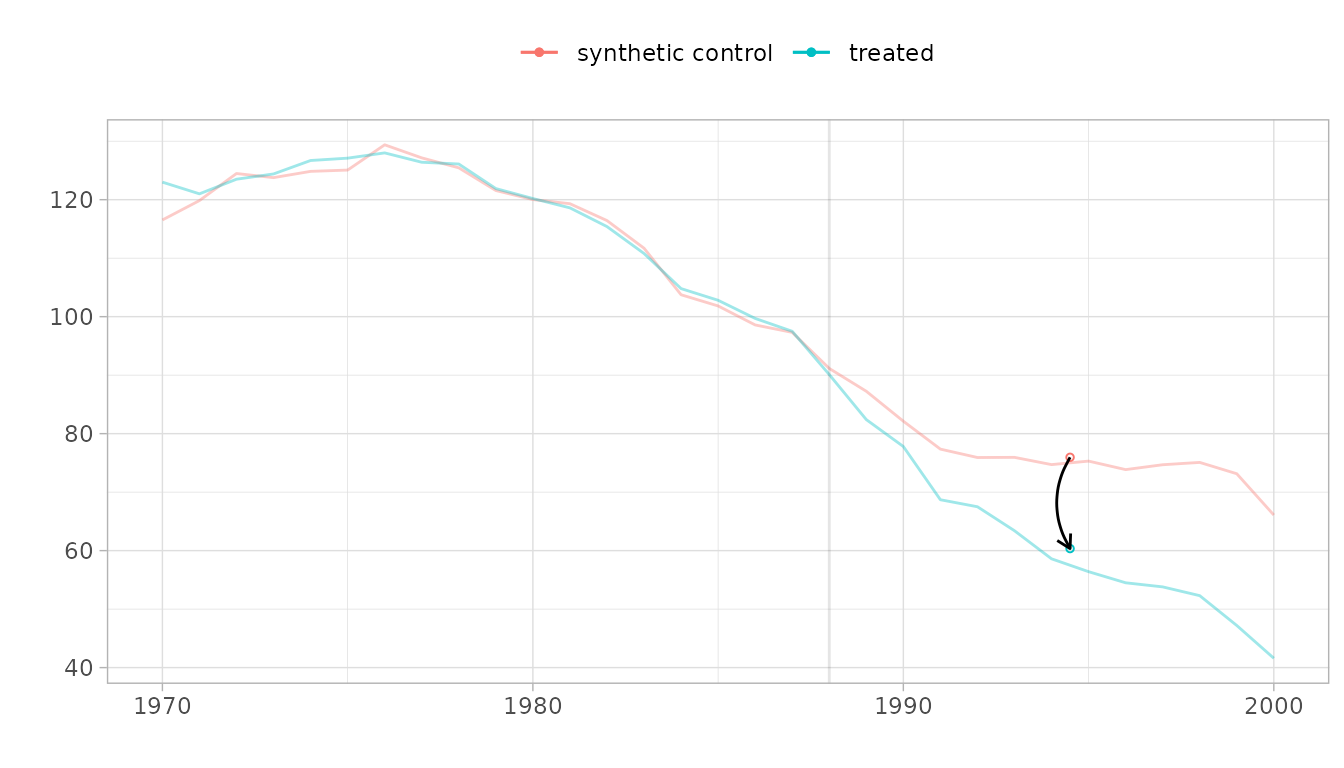
Working with Model Objects
Extracting Components
The new interface creates rich model objects with many components:
# Get the call
attr(result, "call")
#> synthdid(formula = PacksPerCapita ~ treated, data = california_prop99,
#> index = c("State", "Year"))
# Get the formula
attr(result, "formula")
#> PacksPerCapita ~ treated
# Get panel index
attr(result, "index")
#> unit time
#> "State" "Year"
# Get weights
weights <- attr(result, "weights")
names(weights)
#> [1] "omega" "lambda" "lambda.vals" "vals" "omega.vals"
# Lambda (time weights)
head(weights$lambda)
#> [,1]
#> [1,] 0
#> [2,] 0
#> [3,] 0
#> [4,] 0
#> [5,] 0
#> [6,] 0
# Omega (unit weights)
head(weights$omega)
#> [,1]
#> [1,] 0.000000000
#> [2,] 0.003441829
#> [3,] 0.057512787
#> [4,] 0.078287288
#> [5,] 0.070368123
#> [6,] 0.001588411Model Information
# Data information
data_info <- attr(result, "data_info")
data_info
#> $outcome
#> [1] "PacksPerCapita"
#>
#> $treatment
#> [1] "treated"
#>
#> $covariates
#> character(0)
#>
#> $n_units
#> [1] 39
#>
#> $n_periods
#> [1] 31Advanced Usage
Custom Options
You can pass any option from synthdid_estimate() to the
formula interface:
Summary
Quick Reference
| Task | Command |
|---|---|
| Basic estimation | synthdid(outcome ~ treatment, data, index) |
| With SE | synthdid(..., se = TRUE, se_method = "bootstrap") |
| With covariates | synthdid(outcome ~ treatment \| cov1 + cov2, ...) |
| Method selection | synthdid(..., method = "synthdid"/"did"/"sc") |
| Extract coefficient | coef(result) |
| Confidence interval | confint(result) |
| Summary | summary(result) |
| Effect curve | predict(result, type = "effect") |
| Counterfactual | predict(result, type = "counterfactual") |
| Residuals | residuals(result, type = "control") |
| Fitted values | fitted(result) |
| Update model | update(result, method = "did") |
| Plot | plot(result) |
Migration from Old Interface
| Old Interface | New Interface |
|---|---|
setup <- panel.matrices(data) |
— |
synthdid_estimate(setup$Y, setup$N0, setup$T0) |
synthdid(outcome ~ treatment, data, index) |
c(result) |
coef(result) |
sqrt(vcov(result)) |
confint(result) (automatic SE) |
| Manual effect curve calculation | predict(result, type = "effect") |
| Custom comparison code | update(result, method = "did") |
Next Steps
- See
vignette("synthdid")for the original introduction - See
vignette("more-plotting")for plotting examples - See
?synthdidfor complete function documentation - See
INTERFACE_IMPROVEMENTS.mdfor implementation details