Synthetic Difference-in-Differences Estimation with Formula Interface
Source:R/interface.R
synthdid.RdThis function provides a formula-based interface to synthdid estimation,
similar to lm(), plm(), and glm(). It automatically
handles panel data conversion and returns a rich model object compatible
with standard R modeling tools.
Arguments
- formula
A formula of the form
outcome ~ treatmentoroutcome ~ treatment | covariates. The treatment variable should be a binary indicator (0/1).- data
A data.frame containing the panel data in long format.
- index
A character vector of length 2 specifying the names of the unit and time variables, e.g.,
c("state", "year"). If NULL, assumes the first two columns are unit and time.- method
Estimation method: "synthdid" (default), "sc" (synthetic control), or "did" (difference-in-differences).
- se
Logical. If TRUE, compute standard errors. Default is FALSE.
- se_method
Standard error method: "bootstrap" (default), "jackknife", or "placebo".
- se_replications
Number of replications for bootstrap/placebo standard errors.
- ...
Additional arguments passed to
synthdid_estimate().
Value
An object of class c("synthdid", "synthdid_estimate") with components:
- coefficients
Treatment effect estimate
- call
The matched call
- formula
The formula used
- terms
The terms object from the formula
- model
The model frame (if requested)
- Y
The outcome matrix
- N0
Number of control units
- T0
Number of pre-treatment periods
- weights
List with lambda, omega, and beta weights
- setup
List describing the problem
- estimator
Name of estimator used
- index
Panel index variables
- data_info
Information about the original data
Examples
# \donttest{
data(california_prop99)
# Basic usage
result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year")
)
# With standard errors
result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
se = TRUE,
se_method = "bootstrap"
)
#> Warning: bootstrap standard errors require more than one treated unit.
# Standard R methods work
print(result)
#> Synthetic Difference-in-Differences Estimate
#>
#> Call:
#> synthdid(formula = PacksPerCapita ~ treated, data = california_prop99,
#> index = c("State", "Year"), se = TRUE, se_method = "bootstrap")
#>
#> Treatment Effect: -15.6
#>
#> Units: 38 control, 1 treated
#> Time Periods: 19 pre-treatment, 12 post-treatment
#>
#> Convergence: NOT CONVERGED - lambda (10000/10000 iters), omega (10000/10000 iters)
#> Consider increasing max.iter. Use summary() for details.
summary(result)
#> Call:
#> synthdid(formula = PacksPerCapita ~ treated, data = california_prop99,
#> index = c("State", "Year"), se = TRUE, se_method = "bootstrap")
#>
#> Treatment Effect Estimate:
#> Estimate Std. Error t value Pr(>|t|)
#> treated -15.6 NA NA NA
#>
#> Dimensions:
#> Value
#> Treated units: 1.000
#> Control units: 38.000
#> Effective controls: 16.388
#> Post-treatment periods: 12.000
#> Pre-treatment periods: 19.000
#> Effective periods: 2.784
#>
#> Top Control Units (omega weights):
#> Weight
#> Nevada 0.124
#> New Hampshire 0.105
#> Connecticut 0.078
#> Delaware 0.070
#> Colorado 0.058
#>
#> Top Time Periods (lambda weights):
#> Weight
#> 1988 0.427
#> 1986 0.366
#> 1987 0.206
#>
#> Convergence Status:
#> Overall: NOT CONVERGED
#> Lambda: ✗ (10000/10000 iterations, 100.0% utilization)
#> Omega: ✗ (10000/10000 iterations, 100.0% utilization)
#>
#> Recommendation: Consider increasing max.iter or relaxing min.decrease.
#> Use synthdid_convergence_info() for detailed diagnostics.
coef(result)
#> treated
#> -15.60379
confint(result)
#> Warning: Standard error not available; cannot compute confidence interval
#> Lower Upper
#> treated NA NA
plot(result)
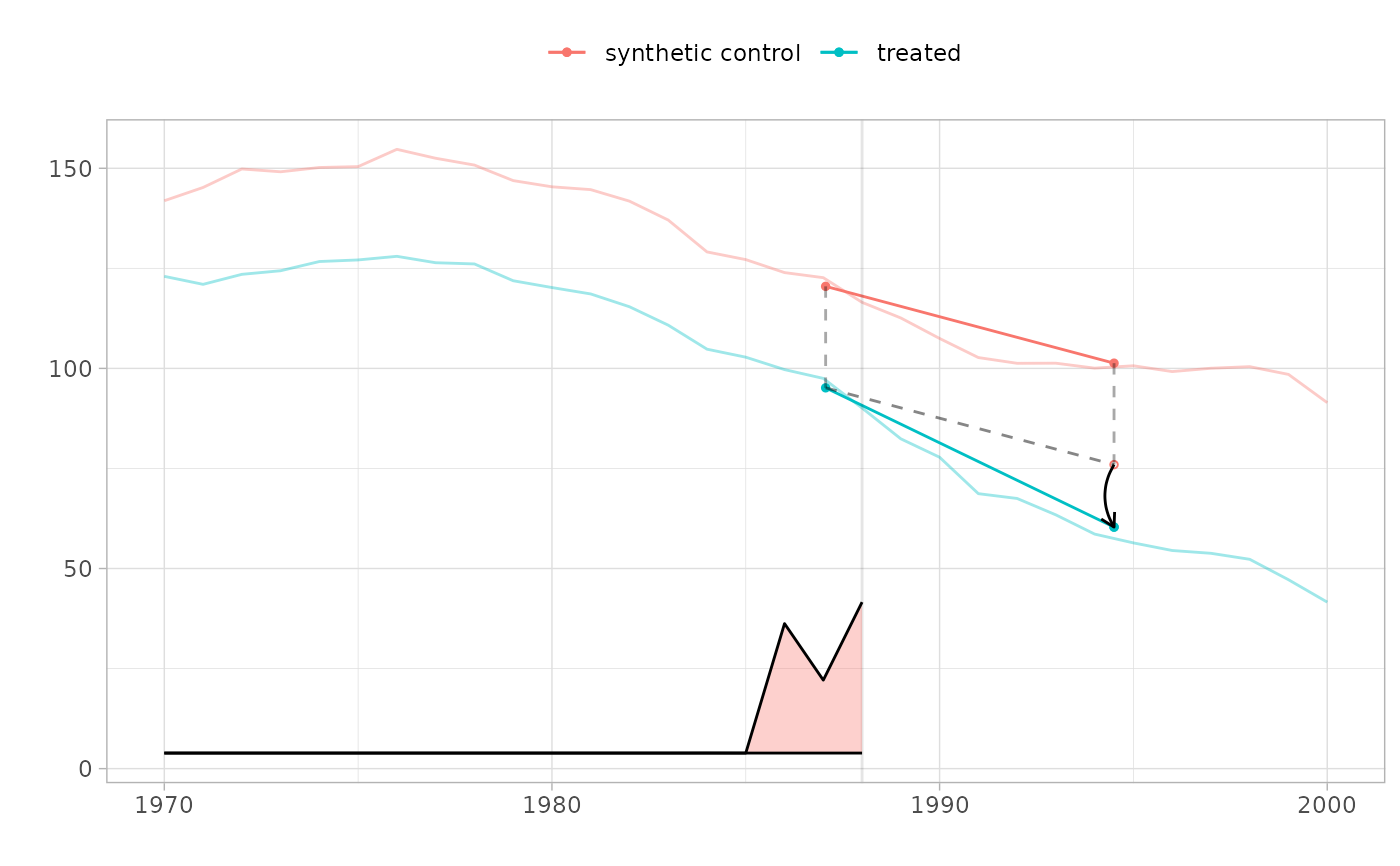 # Compare methods
did_result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
method = "did"
)
# }
# Compare methods
did_result <- synthdid(PacksPerCapita ~ treated,
data = california_prop99,
index = c("State", "Year"),
method = "did"
)
# }